You can use InfraNodus to visualize the discourse in any domain of science using our Scientific Papers Import app. This was a much-requested feature: most of our users wanted to get the data from Google Scholar, which we could not provide because of Google's restrictions. However, we now started to explore other academic APIs and we've already implemented
1) Core.Ac.Uk, which claims to have the largest collection of open access papers in the world;
2) PubMed, which is the main source for medical and life science research.
3) PLOS, which is the leading publisher of open access papers (mainly in the field of life sciences and biology)
4) Google Scholar search results (note, in this case you're visualizing the titles of the papers and the excerpts selected by Google Scholar only, not the whole papers or their abstracts).
If you don't find the source you need, you could also use another service to export the titles and / or abstracts of the papers you're interested in as a CSV file and then import the file using the CSV import app in InfraNodus.
Now you can choose the Scientific Papers Import App, then type in the search query (e.g. "text network analysis") and visualize the discourse that happens around this topic in the scientific community. Here's how you can do this:
Step 1: Open the Scientific Papers Import App from the InfraNodus Apps page.
Note, if you would like to import data from Google Scholar, we recommend you to choose the Google Scholar app:
Step 2: On the actual Import panel you can type in your search query (best results are in English). You can use PubMed (medical and life science papers) or Core.Ac.Uk (open access papers) as the sources, as well as PLOS for now, but we're planning on adding more (let us know which sources you'd like to have).
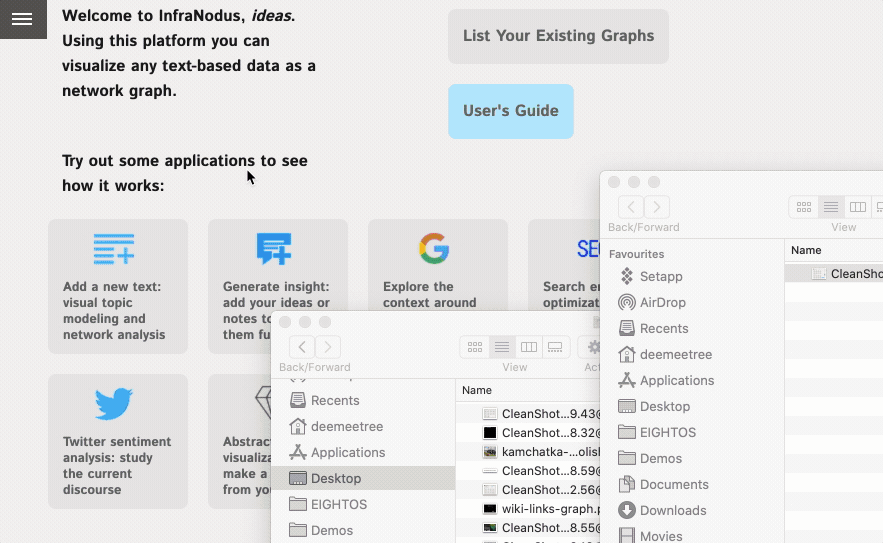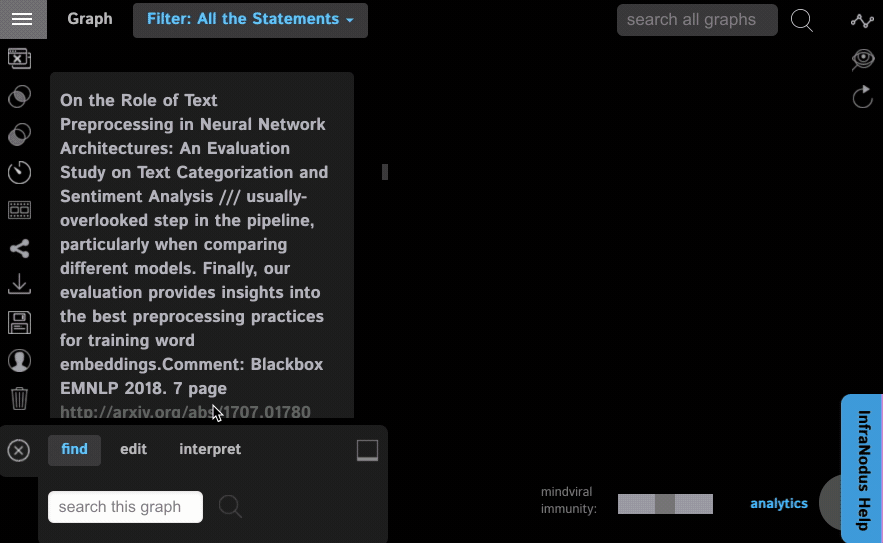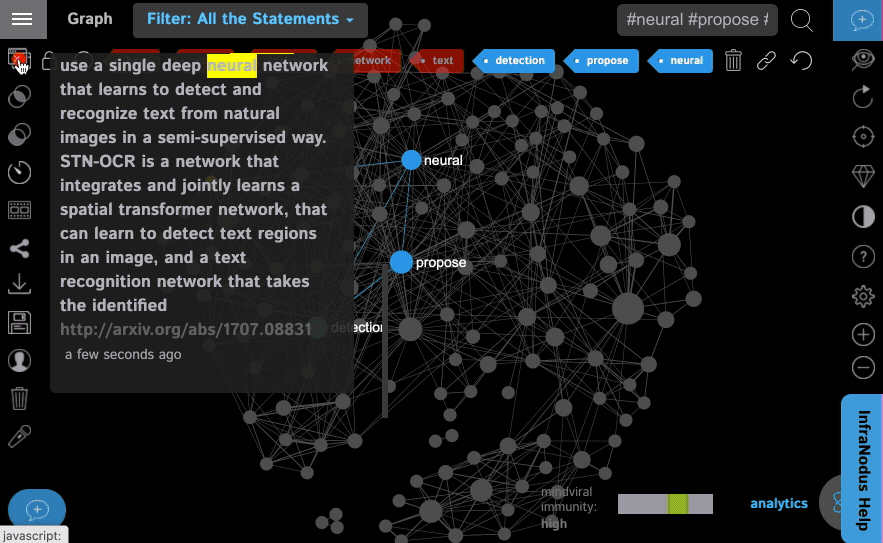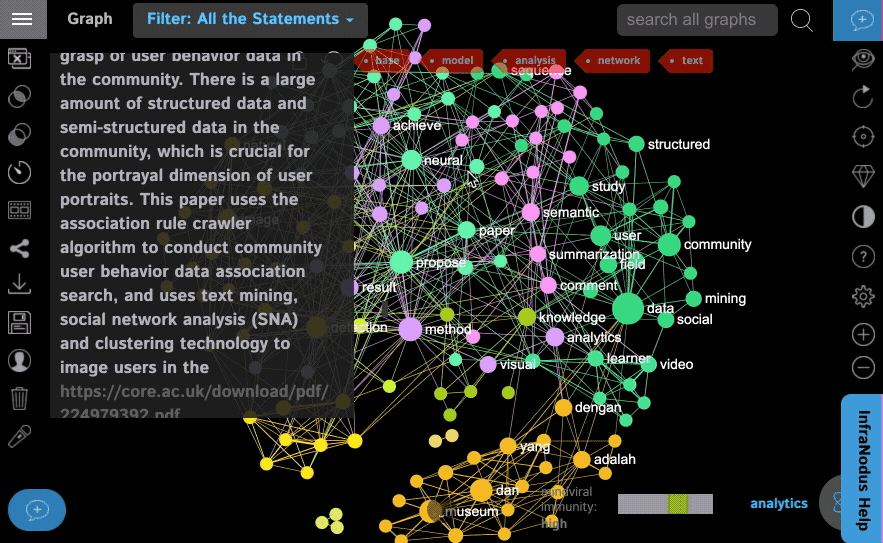
Step 3: Select whether you want to exclude the search terms from the graph (you can always get them in after the graph is loaded). We leave this feature ON by default, because if you turn it off, the search terms you use will be the main terms in the graph and you actually want to see the context around them, so it's better to remove them from the graph by default.
Step 4: Select whether you want to exclude the abstracts. We leave this option OFF by default, because the Abstracts provide quite interesting details on methodology. However, if you just want a general overview of the field, you can turn this off and then only the papers' titles will be included into the graph. Which is also interesting, because it lets you
Step 5: Select how many papers you want to extract. Note, that the system will extract the most relevant papers sorted by relevance by Core.Ac.Uk API — whatever their system thinks suits your search term the best. We set the default value of papers to extract as 40 as this is generally 4 pages of search results and also provides a nice readable graph with not too much irrelevant data. However, you can select up to 100 papers if you're looking for additional insights (maybe on your 2nd try).
Step 6: Choose the name for your graph. We recommend to use lowerdash _ instead of spaces and prefix this type of graph with "science_" so you know what you're looking at.
Step 7: Click Import and visualize the results. The Analytics panel will show you the main topics that exist in the scientific discourse for this particular topic. Note, that the search terms you used will be hidden from the graph (at the top right) so you can see what's hiding behind them. You can always get them back into the graph if you click on them.
Methodology: our algorithm visualizes the abstracts and the titles of the open access papers found for your search query. The words are represented as the nodes and their co-occurrences are the connections between them. Based on community detection algorithms and force-atlas layout we identify the groups of nodes that are more densely connected together than with the rest of the network — these form so-called topical clusters. The nodes / words that have the highest betweenness centrality measure are shown bigger on the graph — having the highest influence. These are used to indicate the top connecting terms. Read more on the methodology in our peer-reviewed white-paper presented at WWW 19.
Step 8: Once you visualize the graph, start by deleting the obvious terms from the graph (you can always get them back in), so you can get to the interesting topics. You can also select the most interesting nodes on the graph and then see the relevant papers / abstracts that have the highest concentration of these terms, along with the links to the actual research papers.
Step 9: Using the Analytics > Insight panel you can reveal the structural gap in the discourse: indicating the two topics that could be connected better, which may, in turn point you to an interesting research question that you could ask in relation to the discourse around your topic. This is also available in the Filter: Statements to Link menu at the top left.
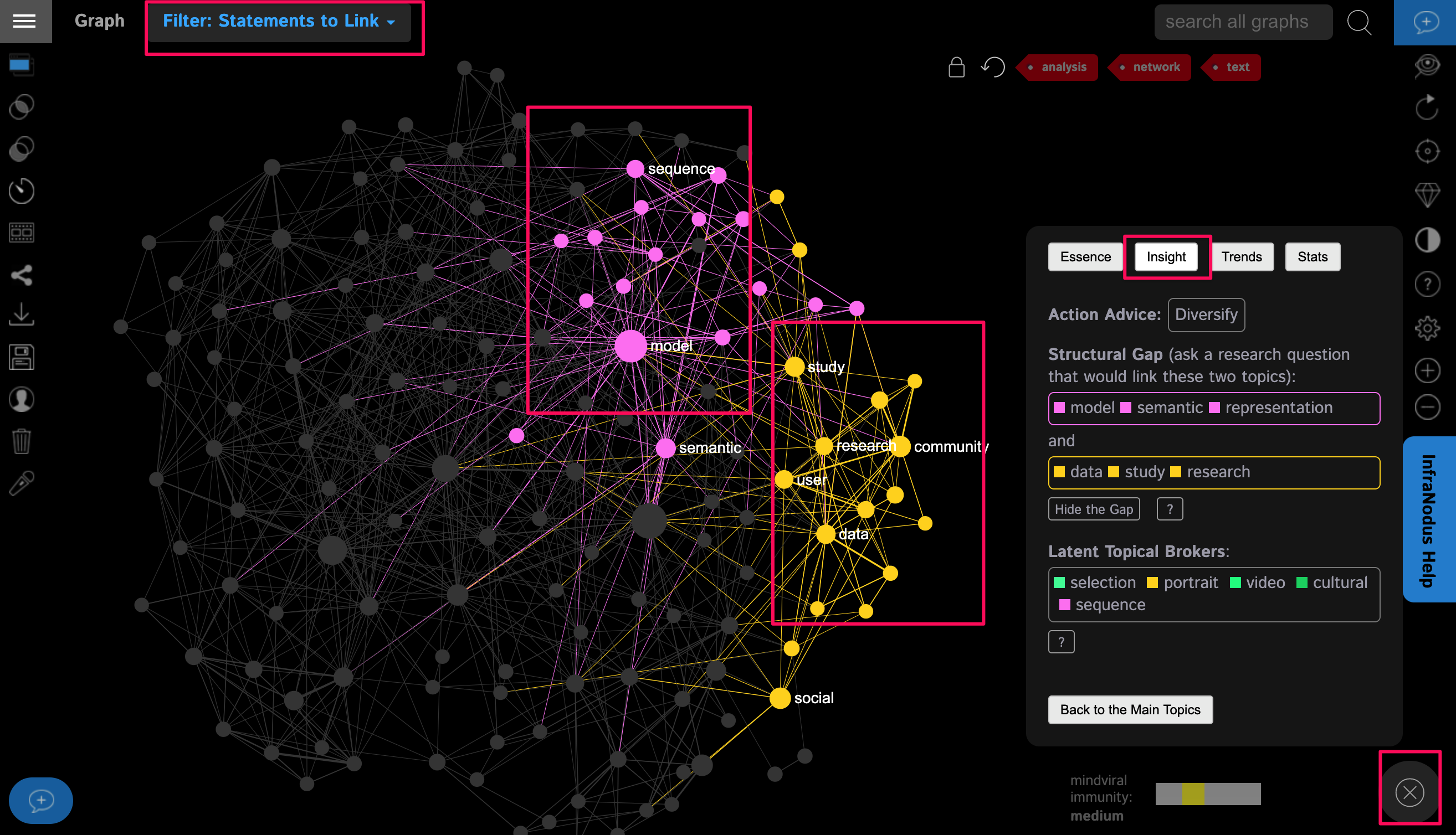
Step 10: We encourage you to choose a few concepts or topics on the graph, and to click the nodes to see in which context they're used and the scientific papers that contain these concepts. This is a very useful way to go deeper into a specific topic and to explore the nuances.
Step 11: If you find a tangent you like (for instance, you were searching for "deep sleep" and then found that exercise has some benefits for "deep sleep") you can import more data on this topic. In order to do that, just choose + Add / Import, then click the import button and choose your source.
If you import the data into the same graph using Google Scholar, your new results will have the new search query as a tag, so you can filter them after:
If you imported data into a different graph, you can combine them after using the graph comparison feature.
Step 12: Reiterate

Comments
0 comments
Please sign in to leave a comment.